A team of scientists working at beamline 9.0.1 of the Advanced Light Source (ALS) at the U.S. Department of Energy's Lawrence Berkeley National Laboratory has used x-ray diffraction microscopy to make images of whole yeast cells, achieving the highest resolution-11 to 13 nanometers (billionths of a meter)-ever obtained with this method for biological specimens. Their success indicates that full 3-D tomography of whole cells at equivalent resolution should soon be possible.
"We have demonstrated that lensless imaging techniques can achieve very high resolution while overcoming the limitations of x-ray optics-limitations that include requiring 20 to 50 times the radiation exposure to get a magnified image of the sample," says Chris Jacobsen, formerly of Stony Brook University, now of Argonne National Laboratory and Northwestern University, who designed the lensless-imaging research program at beamline 9.0.1. "While at present it takes us a long time to image a single specimen-and full 3-D imaging of hydrated cells will take even more work-this is a big step in the right direction."
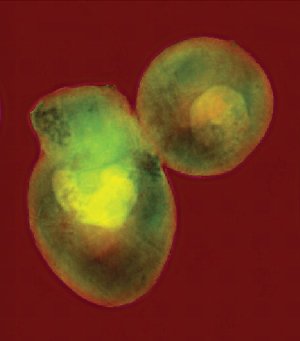 A pair of yeast cells imaged at very high resolution using coherent soft x-rays at the Advanced Light Source’s beamline 9.0.1. The coherent (laser-like) beam of penetrating x-rays allows a computer to reconstruct the cells’ internal structures from a diffraction pattern, without focusing the light with a lens.
A pair of yeast cells imaged at very high resolution using coherent soft x-rays at the Advanced Light Source’s beamline 9.0.1. The coherent (laser-like) beam of penetrating x-rays allows a computer to reconstruct the cells’ internal structures from a diffraction pattern, without focusing the light with a lens.
Three-dimensional imaging of whole cells under conditions close to those in nature, namely a hydrated (watery) environment, is already done at the National Center for X-Ray Tomography at ALS beamline 2.1, under the direction of Carolyn Larabell of Berkeley Lab's Physical Biosciences Division, where large numbers of cells can be processed in a short time at resolutions of 40 to 60 nanometers. The ability to increase resolution to the 10-nanometer range would significantly advance research in both biology and materials sciences.
"Ten-nanometer resolution is easy to achieve with an electron microscope," says Janos Kirz of the ALS, co-designer with Jacobsen of the lensless imaging program. "The problem is that electron microscopy is limited to very thin samples, a few hundred nanometers or less-so you can't use it to look through a whole cell."
While x-rays have the ability to look deep into thick specimens, or right through them, imaging with a lens has its own problems. Even the best x-ray microscope lenses (concentric circles of metal known as Fresnel zone plates), can't focus x-rays with high efficiency, so to get an image means using such intense radiation that it more quickly damages biological specimens. At the same time, the geometry of the highest-resolution zone plates makes for an extremely narrow depth of focus.
To get around these barriers, a research team led by Jacobsen's students Johanna Nelson, Xiaojing Huang, and Jan Steinbrener-also of Stony Brook-used lensless x-ray diffraction microscopy. To produce a high-resolution diffraction pattern from noncrystalline structures like the membranes and organelles of a cell, the light has to be coherent, that is, laser-like, having all the same frequency and phase. Beamline 9.0.1 was built to supply this kind of light.
As they proceed through the cell, the coherent x-rays are scattered and differentially absorbed by the cell's internal structures. There's no lens either in front or behind the sample as the light passes through the cell and reaches the detector, so there's nothing to limit resolution or efficiency.
But the result looks nothing like an image. Instead it's a pattern of dark and light speckles, the traces of the scattered x-rays. A computer-which acts as the "lens" in lensless imaging-uses these patterns to create an image.
Stefano Marchesini, ALS beamline scientist for beamline 9.0.1, says, "The challenge of the lensless technique is that essentially the preparation and quality of the sample have to be perfect-and, ideally, completely isolated in the beam."
The ideal can't be reached in reality, since the sample has to be supported. Furthermore, to image hydrated cells, the specimen has to be frozen, which introduces misleading data from the presence of ice. Last November Jacobsen's team used beamline 9.0.1 to image a frozen, fully hydrated yeast cell at a resolution of 25 nanometers-a resolution limited by ice. The current experiment's 10 to 13 nanometer resolution required using unhydrated, freeze-dried cells at room temperature.
Until recently, to produce an image, researchers had to know the precise shape of the sample's support and have a pretty good idea of the shape of the sample itself, before the computer could even start solving the diffraction patterns. A new algorithm written by Marchesini called "shrinkwrap" converges on the diffraction data through subsequent iterations, and finally distinguishes and subtracts the support from the sample image.
Nevertheless, tens of thousands of iterations with both algorithms and manual adjustments of the cell boundaries were needed to yield the current experiment's final image of a pair of yeast cells. Details emerged when the computer was instructed to represent differences in absorption with differences in image brightness, and differences in the phase of the light, altered by scattering, with differences in image color.
The relationship of a cell's internal structures can only be determined accurately by full 3-D tomography. In a 2-D image these features are stacked one on top of another, and separating the planes is partly a matter of guesswork. Short of true 3-D, it's possible to gain some sense of how the internal structures are arranged by focusing on different depths in the structure, then comparing these images with others of the same object made with different techniques.
In the current experiment, additional images of the same freeze-dried cells were made first by scanning transmission x-ray microscopy (using a lens), which was of lower resolution but helped confirm features at different planes, and then by scanning electron microscopy, which was of higher resolution but could only show surface details such as sugar molecules in the cell walls, which the researchers had labeled with gold particles to serve as position markers.
"True 3-D of whole, hydrated, frozen cells at this very high resolution is our next step," says Jacobsen. "One ingredient is to rotate the frozen sample in the beam to get more depth information. Another is to scan the beam across the sample."
Jacobsen's team is making progress using both methods at beamline 9.0.1, but the present set-up, remarkable as it is, requires what he calls "heroic" experimental efforts. To reap the full benefits of lensless imaging with x-ray diffraction microscopy, he says, will require a source of intense coherent light that can yield a thousand times the current data rate. "We'll want to readily tune the frequency of the light, and tune its energy to the optimum. And we'll need a sample stage that can easily tilt and rotate frozen samples, holding them in the beam for many exposures. These advances would mean we could do our experiments reasonably rather than heroically, and on a wider range of materials."
In the meantime, x-ray diffraction microscopy at beamline 9.0.1 is pointing the way to what lensless imaging can do.
This work was supported by the U.S. Department of Energy's Office of Science and the National Institutes for Health.