Atomic force microscopy (AFM) has the potential to disclose several properties regarding a material. Most recently, it has been used to achieve topographical information, but it can also probe electrical conductance, mechanical stiffness, magnetism, and resistivity.
Scientists have utilized it to study interactions between enzymes and their substrates,1 structural changes in injured or diseased tissue,2 macromolecular interactions between lipids3 and analysis of nucleic acid organization and structure,4 to name some applications.
AFM executes analyses on a micro and nanoscale, thereby enabling it to measure phenomena as minuscule as electrostatic interactions, van der Waals forces, and molecular bonds.5 AFM also has the potential to produce high-resolution, elaborate images of sample surfaces. This exhibits micro and nanoscale properties of materials as flat as cleaved mica or as non-uniform as a cell.
A fascinating feature of AFM is its potential to quantify multiple micro-and nanoscale properties in a single test on samples that seem to be unstained, unfixed, and alive. Of particular use in many fields is the simultaneous measurement of mechanical properties and topographical features.
Conventional light microscopy has the potential to disclose a wealth of information regarding a sample, particularly a biological one. Light microscopy can give information to the investigators on the shape of a cell, localization of subcellular structures inside the cell, and even organization of cellular infrastructure, among several other parameters.
However, a limitation with such optical data is that it is unable to quantify, in a direct quantifiable way, the mechanical properties of that cell; such properties provide investigators with significant information regarding the cytoskeletal organization and phenotype of the cell.
The stiffness of the cell, evaluated by measuring the elastic modulus of the cell, is different at several points throughout its surface; cells tend to be stiffer over cytoskeletal structures and softer over the cytoplasm. In general, AFM can evaluate both mechanical and topographical properties of any material, such as cells, concurrently in a single assay.
Since the 1990s, AFM scientists have utilized force mapping to concurrently offer nanoscale mechanical and topographical data regarding a substrate. Force mapping involves producing separate force curves at distinct points on a material, which are further utilized to assess height values and stiffness.6,7
This workhorse method is easy and direct, thereby making it simple to implement with any AFM setup. But it can also be explained as a comparatively slow procedure with low lateral resolution - which is not ideal for several biological applications.
A novel method referred to as “force scanning,” was developed by the researchers to produce both structural and functional data regarding the material of interest via a much more rapid process8 (Figure 1).
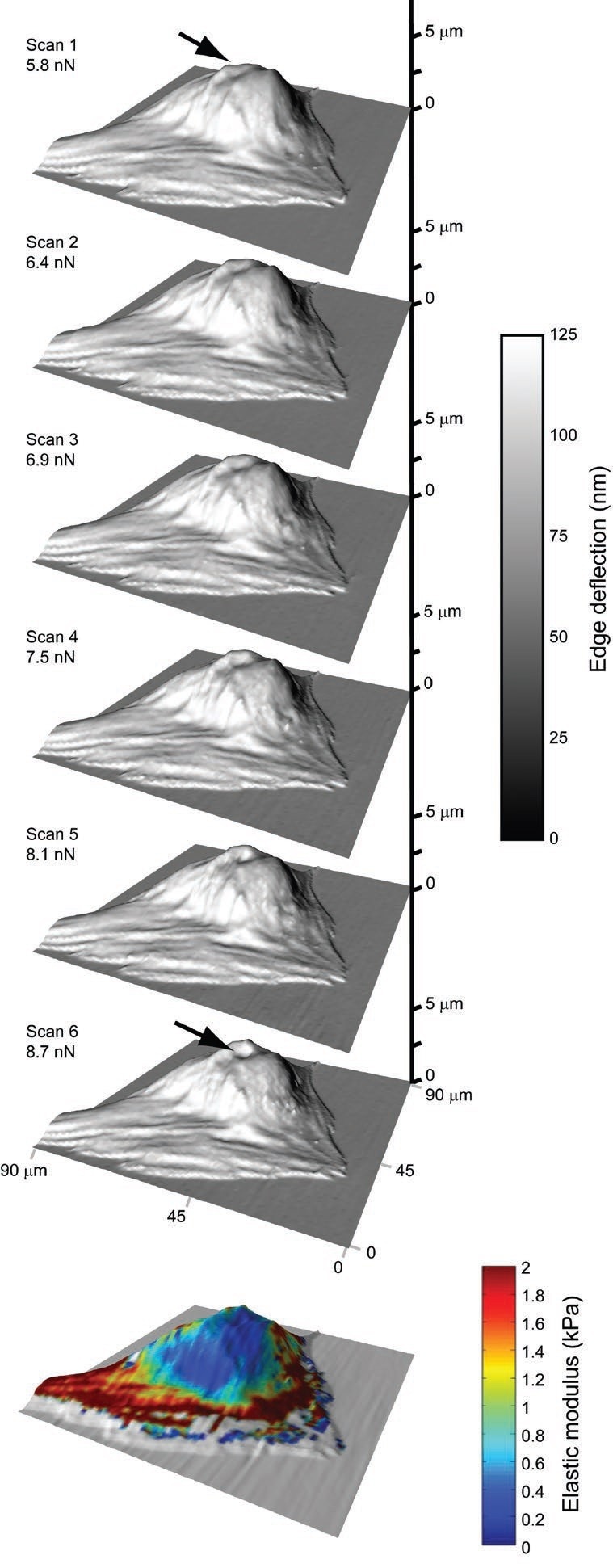
Figure 1. An incremental series of quick, contact-mode scans can be stacked to capture force-indentation curves for all points in an image. Elastic moduli can then be extrapolated from these data, producing high-resolution, spatial modulus maps. The arrow points to a region of greater compliancy located over the cell nucleus. Image Credit: Asylum Research - An Oxford Instruments Company
This AFM-based application has the potential to quickly capture a substrate’s high-resolution topographical images (a resolution of 32×32 points takes just seconds when utilizing scanning frequencies ranging between 2 to 5 Hz) which are further utilized to measure location-specific, mechanical properties at the time of post-processing.
Force scanning is an easy method to perform and it is an extensively applicable method to examine compliant materials. The method can be utilized for any deformable material, ranging from agarose gels to cartilage to living cells.
It has been displayed to accurately map gel surfaces, measure cell-cell interactions, and specify living cells and extracellular matrices at high resolutions,7 which is an enhancement from earlier attempts.6
Force scanning does not require unique cantilevers, equipment, or alterations to be made to standard AFM technology. So long as a clear contact-mode image could be achieved, force scanning is a feasible option.
For the versatility of force scanning to be illustrated, a range of samples were evaluated with the help of the Asylum Research MFP-3D-BIO™.
Atomic Force Microscope, which is attached to an inverted optical microscope. Cantilevers were altered with 5 µm borosilicate glass beads to guarantee well-defined, conformal contact along with the sample surface.
Indentation with a spherical indenter enables modeling of the mechanical properties of the substrate through the Hertz contact model, which states that when a spherical indenter presses into an elastic surface, the surface area of contact raises and the applied force can be linked to the displacement created by the indenter.
Spring constants were evaluated from the power spectral density of thermal noise fluctuations before each experiment.9 For data collection to be performed, an Asylum Research function, “LongTermScanning.ipf” (available through the AR Forum), was altered to quickly gather a series of contact-mode scans.
Originally, this function was developed to gather several high-resolution adjacent scans, and stitch them collectively to make a single high-resolution scan that has the potential to adjust voltage drift over time.
For the force scanning application, this function was changed so that a single region was scanned in a repeated manner but with incrementally greater setpoints to make a stack of topographical images. Scans were performed at a 90° angle, thereby recording z-piezo movement and normal or lateral deflections.
While making use of a 0 °scan angle is an option, this can increase the chance of sample damage and hence makes it hard to consider frictional forces. Then, this can be quantified with the help of data gathered from the lateral deflection of the cantilever.
But for the examples shown in this article, lateral forces were discovered to have little effect on the quantified mechanical properties since the employed and normal forces were much greater in magnitude, thereby swamping any lateral contributions effectively.
Although not explored in this article, it would be theoretically possible to select particular testing parameters (for example, scan rate, the normal or lateral stiffness of the cantilever, setpoint force, tip geometry) that enable simultaneous measurement of frictional properties, material stiffness, and surface topography.
Irrespective of the method, all recorded data has been post-processed and examined with the help of custom MATLAB scripts, thereby integrating a Hertz contact model. Such scripts correct for any recorded drifts at the time of the test (for example, z-piezo or laser) and assess efficient indentation forces from the lateral and normal cantilever deflections.
Correct calibration of the cantilever seems to be essential to transforming voltage signals into applied force. It must be observed that the contact mechanics included in this article are very complicated, and it has been suggested that a control material of familiar stiffness be utilized to confirm the precision of the procedure for a particular equipment setup.
Force scanning itself depends on the AFM executing a raster scan throughout a surface while retaining an (almost) constant applied downward force (that is, setpoint). The force makes the cantilever notch into the surface while scanning is being done and thereby efficiently tracing the material of interest and producing a precise topographical map, albeit slightly deformed in the vertical direction as a result of the setpoint force.
Multiple scans are performed with incrementally increasing setpoint forces, indenting the surface more and more with each iteration (Figure 2). Usually, 5–7 scans are performed per sample.
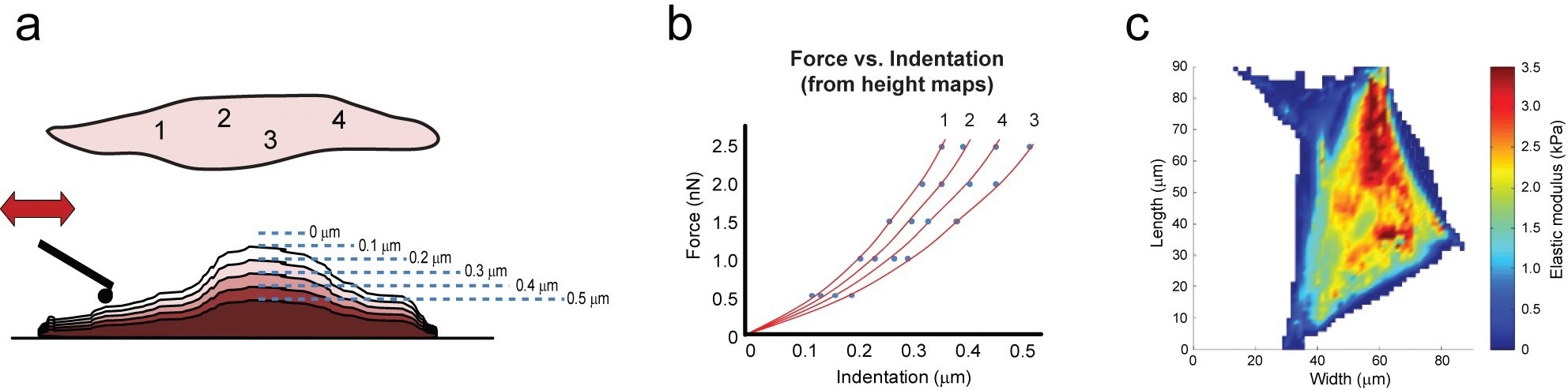
Figure 2. Force scanning involves combining a series of contact-mode scans that deform a material with incrementally larger setpoint forces (a). Using height and setpoint data, force vs. indentation curves can be constructed for all points across a sample, which are then fit with mathematical models to obtain mechanical properties (b). Overlaying this information on the sample can reveal structure-property relationships (c). Image Credit: Asylum Research - An Oxford Instruments Company
Correct setpoint values can alter depending on the indenter geometry (for example, 5 µm sphere) and sample characteristics (stiff versus. soft). Here, both must be identified before force scanning. A few indentation curves caused on the sample must be enough to identify a suitable range of values (for example, 4 to 12 nN for agarose, 150–500 nN for cartilage, 0.5–4.5 nN for cells,).
The amount of indentation and deflection will change with the stiffness of the substrate. Hence, the setpoint range maxima and minima are identified by executing single force indentation curves caused on every material.
A setpoint that is set extremely low will not produce constant deflection of the cantilever, while a setpoint that is too high will breach the mathematical model or harm a living sample. Each scan offers raw data for normal deflection, heights, and lateral deflection, which could be transformed simply into applied indentation and force.
The relationship present between these two values explains the material’s elastic modulus. Direct comparisons of force scanning and force mapping are hard to carry out as force scanning does not employ discrete force curves at particular points on a substrate but rather produces them from the data.
But outcomes for the two techniques on the same sample are exceptionally similar; validation tests on agarose gels displayed that quantified elastic moduli did not alter considerably (Figure 3).
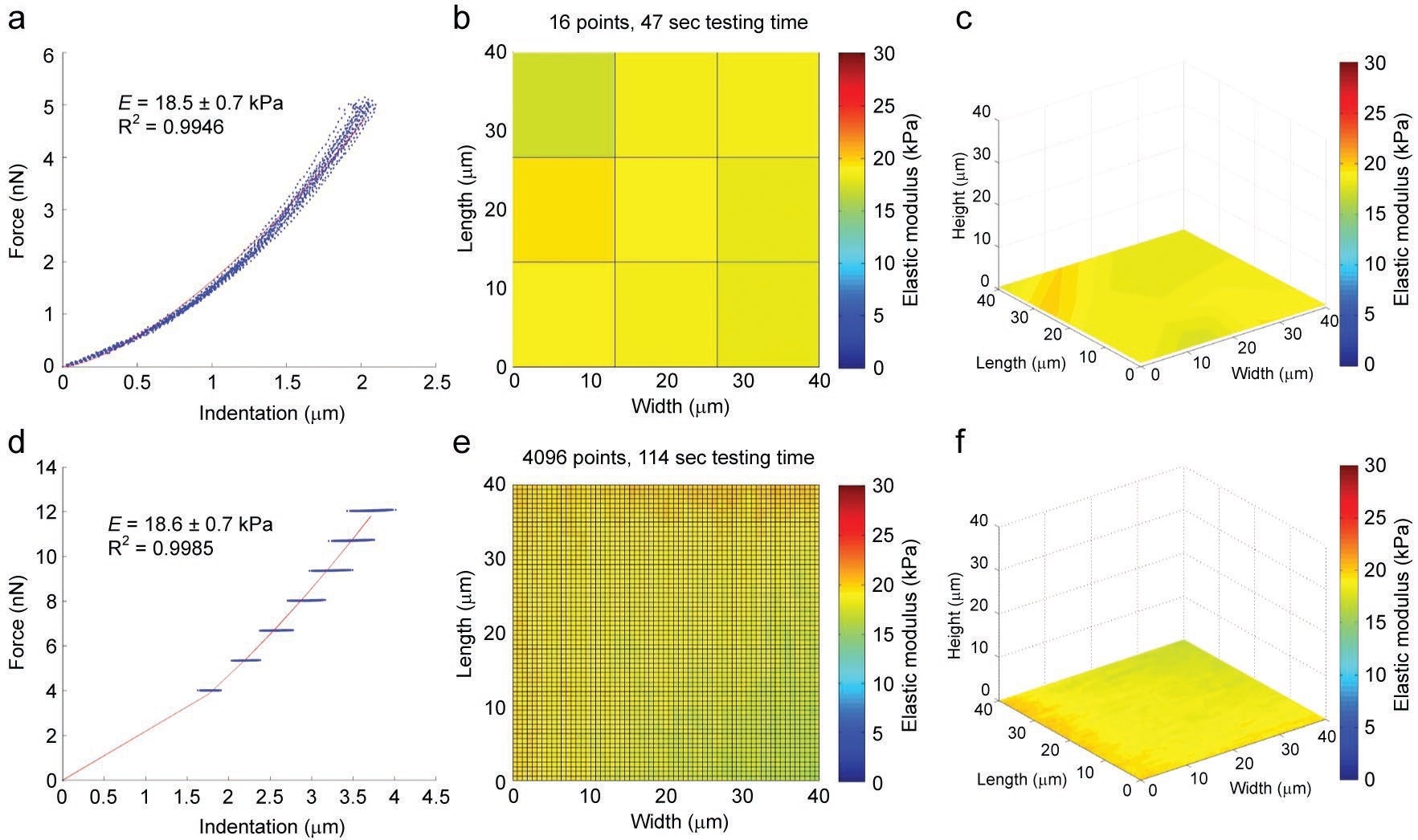
Figure 3. Validation of the force scanning technique was done in part using a soft agarose gel. The standard approach, force mapping, involves applying a series of single indentations to obtain mechanical and topographical data, although at relatively low lateral resolution (a, b, c). Force scanning of the same sample can achieve much more detailed images by sacrificing force curve resolution (d, e, f). The mechanical characteristics of the sample are virtually identical. Image Credit: Asylum Research - An Oxford Instruments Company
The significant advantage of utilizing force scanning is the ease and speed with which the tests are done. Surface images could be done by utilizing a range of scan rates that has the potential to shorten or lengthen the entire process.
In this context, the examples presented normally took scans at 1 to 5 Hz (20-–450 µm/second scan rate, based on region size). This offered a good balance between good image quality and short testing times. The other factor that is a part of force scanning duration is the number of setpoints that have been utilized to produce the force curve, with each one needing an entire scan.
At least 4 to 5 are suggested for fitting the Hertz model, but more can be taken if wanted. Simultaneously, force mapping needs time to take not only the separate force curves (normally done at 0.5 to 15 µm/s) but also time to move from point to point.
This process could be prolonged if the comparative height of the sample alters greatly, as full force curves (for example, 5 to 20 µm in length) are essential to retaining tip-sample separation before indentation, and considering any artificial changes caused in the cantilever deflection at the time of the approach.
This converts into a much slower process compared to force scanning. Depending on empirically determined testing times, if a scan size of 2056 × 2056 points is preferred, it would take nearly a year and a half utilizing force mapping, whereas force scanning reduces that time down to only an hour (Figure 4).
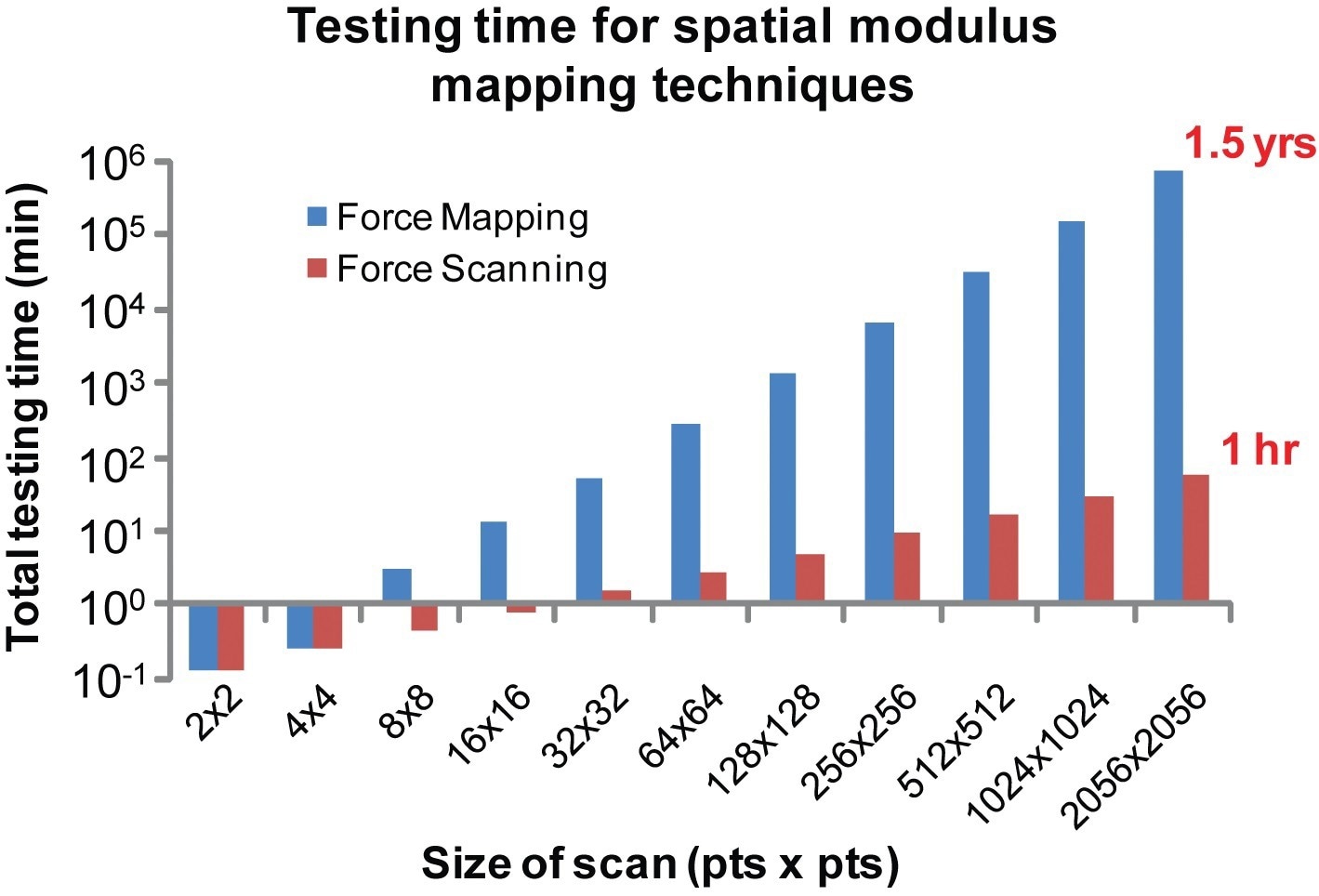
Figure 4. High-resolution scanning takes prohibitively long using standard techniques. Based on temporal trends observed during testing, very high-resolution images could only be achieved using force scanning within a reasonable time frame. For example, a scan size of 2056 × 2056 points would require over a year to complete using force mapping, whereas force scanning cuts that time down to just an hour.8 Image Credit: Asylum Research - An Oxford Instruments Company
While such testing times can change based on the sample characteristics and the scanning parameters used, normally force scanning turns becomes faster by trading off force curve resolution in exchange for speed and entire image resolution. Test duration seems to be significant for many samples and is completely crucial for living materials like cells.
Force scanning is fast enough that tests carried out on living samples have the potential of being completed before major conformational changes take place, an ability that is lacking in conventional force mapping.
Force scanning disclosed a wealth of data when being utilized on biological samples that might not be possible if using a slower method like force mapping. Subcellular features could be differentiated in a topographic and mechanical manner, the nucleus in particular.
The variation in elastic modulus between the cell cytoplasm and the cell nucleus is visible at the time of a force scan, as observed by the change in elastic modulus between the cytoplasmic and perinuclear regions of the cell displayed in Figure 5.
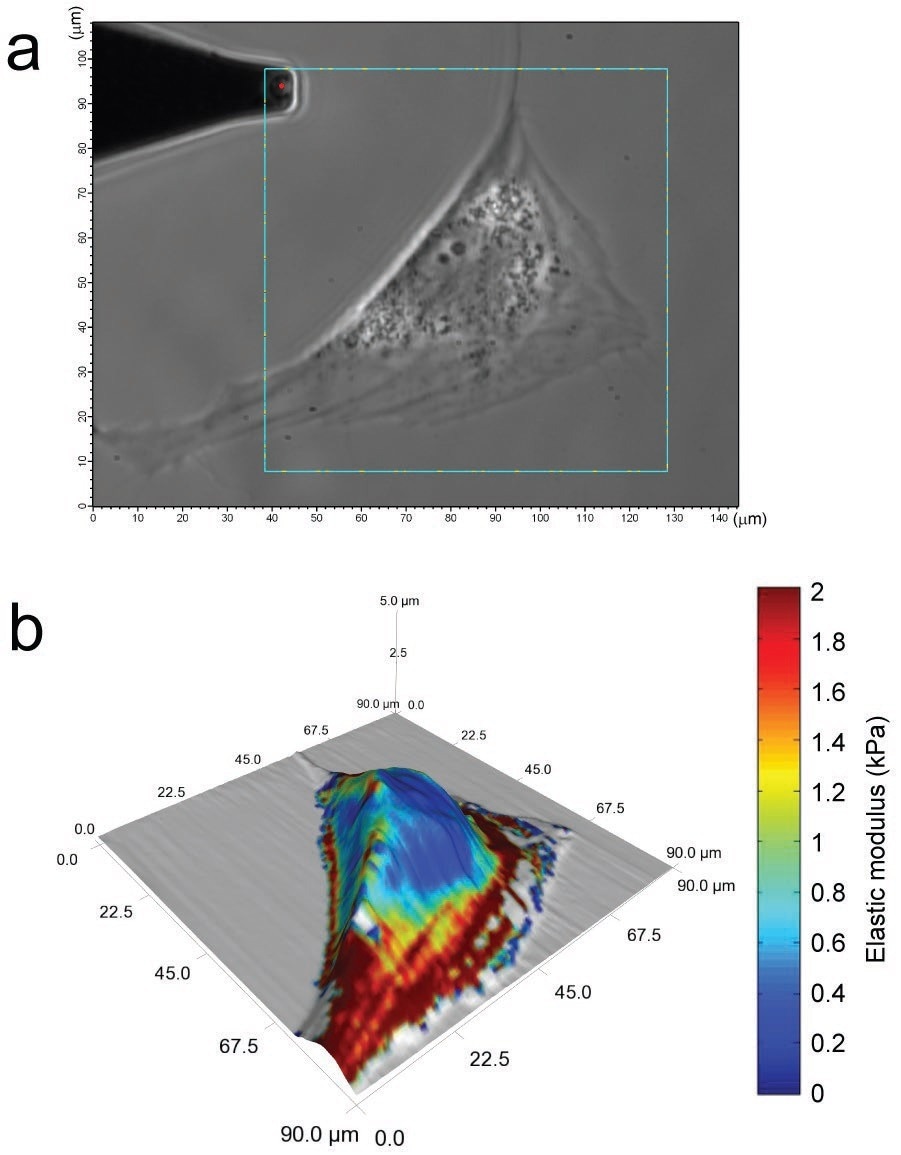
Figure 5. Living, adipose-derived stem cells (ASC), viewed using phase contrast imaging, can be assessed for mechanical property variations across their surface (a). A 128x128 point scan reveals lower moduli in the perinuclear area and higher moduli over cytoskeletal actin filaments (b). This detailed, topographical and mechanical property information was obtained from a six-minute scan.8 Image Credit: Asylum Research - An Oxford Instruments Company
Furthermore, spatial modulus mapping can disclose the mechanical characteristics for lamellipodial extensions and stress fiber distributions, thereby composing an elaborate and informative image of the cell under analysis.
The force scanning method has also been utilized to analyze cell-cell interactions and the mechanical characteristics of biological tissues (Figure 6).
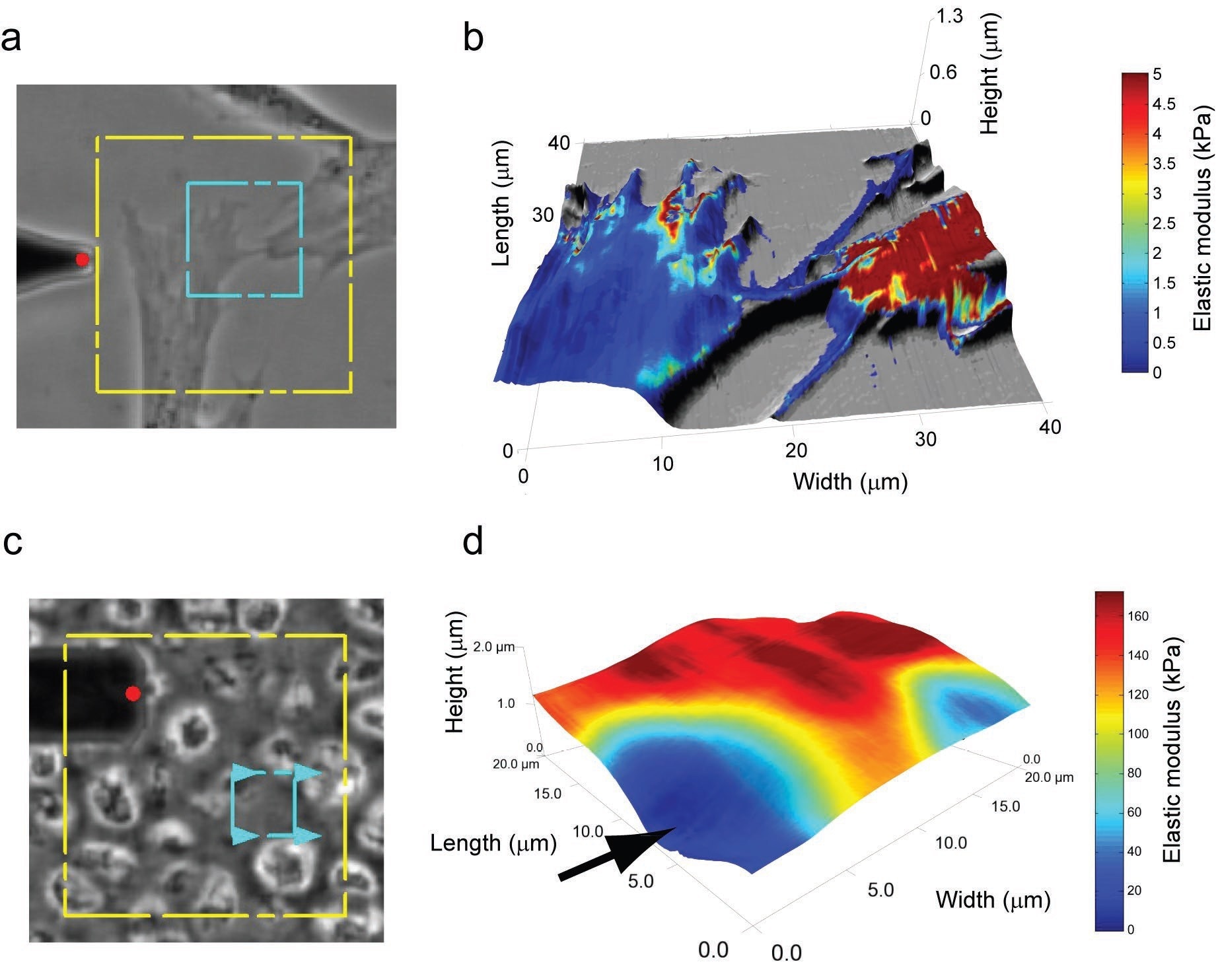
Figure 6. Spatial modulus maps can be used to investigate cell-cell interactions (a-b). Variation in elastic moduli between the two cells is clear, with one cell being much stiffer than the other. Furthermore, filopodial extensions are shown to be much softer than the cell body, although the measurement accuracy for these extremely thin regions is still under investigation. Force scanning also shows excellent applicability for probing the mechanical properties of biological tissues, like the extracellular and pericellular matrices of articular cartilage (c-d). In the rotated force scanning image, the arrow denotes where a cell would be in the native tissue, and the tissue gets progressively stiffer moving outward from the cell edge to the pericellular matrix to the extracellular matrix. The dotted, blue box in (a) and (c) defines the scanned regions. Image Credit: Asylum Research - An Oxford Instruments Company
While the elastic moduli linked to these two materials are orders of magnitude different, the force scanning application can precisely determine the mechanical properties of either by utilizing the correct cantilever and scanning parameters.
Also, force scanning is broadly adaptable and applicable to a range of fields; although it works especially well with biomaterials and soft biological samples, it has applications in several other fields and areas of research as well.
This technique can easily be utilized to quantify the cosmetic and mechanical features of any deformable material that can be evaluated utilizing a contact-mode AFM. Force scanning can be performed in fluid or air environments, on stiff or soft materials, in the absence of modification of standard AFM technologies.
Post-processing of the data is essential for the mechanical properties to be calculated. However, this can be done by making use of a range of available software packages (for example, MATLAB).
Even though this article concentrates on using force scanning to measure mechanical and topographical features, adaptations to the method might set the stage for other possibilities for analysis.
For instance, functionalization of the cantilever tip could provide data on frictional properties or adhesion forces in comparison to data gathered by making use of a non-functionalized tip. In the same way, the frictional properties of a surface could be measured with the help of lateral force data.
Even in the absence of such new adaptations, being able to quickly measure the mechanical characteristics of biological samples in connection to physical structures enables the study of several phenomena.
The so-called mechanotransduction, in which cells tend to translate mechanical signals into biological responses, needs insight into the mechanical properties of the cell and environment. Cell attachment and movement are other processes that require mechanical changes at the subcellular level which could be examined using this method.
Force scanning is an easy, rapid, and easy-to-implement method, thereby setting the stage for higher throughput analyses with higher resolution and detail. With the help of this application, it is possible for scientists to better analyze cell phenotype, cytoskeletal organization, and microenvironment materials under physiological conditions. This enables the study of healthy and living samples.
References
- M. Carrion-Vasquez, A.F. Oberhauser, S.B. Fowler, P.E. Marszalek, S.E. Broedel, J. Clarke and J.M. Fernandez, Mechanical and chemical unfolding of a single protein: A comparison. Proceedings of National Academy of Sciences. 96 3694 (1999).
- A.W. Bridges, N. Singh, K.L. Burns, J.E. Babensee, L.A. Lyon, A.J. Garcia, Reduced Acute Inflammatory Responses to Microgel Conformal Coatings. Biomaterials, 29, 4605–4615 (2008).
- J. Park, Probe Chemistry Effect on Surface Properties of Asymmetric-Phase Lipid Bilayers Colloids and Surfaces B: Biointerfaces 75, 290-293 (2010).
- M. Rief, H. Clausen-Schaumann and H.E. Gaub, Sequence-dependent mechanics of single DNA molecules. Nature Structural Biology 6, 346 (1999).
- J. Vandiver, D. Dean, N. Patel, C. Botelho, S. Best, J.D. Santos, M.A. Lopes, W. Bonfield, C. Ortiz, Silicon Addition to Hydroxyapatite Increases Nanoscale Electrostatic, van der Waals, and Adhesive Interactions Journal of Biomedical Materials Research Part A 78A, 352-363 (2006).
- D.R. Baselt and J.D. Baldeschwieler, Imaging spectroscopy with the atomic-force microscope. Journal of Applied Physics, 76, 33–8 (1994).
- M. Radmacher, J.P. Cleveland, M. Fritz, H.G. Hansma and P.K. Hansma, Mapping interaction forces with the atomic force microscope. Biophysics Journal, 66, 2159–65 (1994).
- E.M. Darling, Force scanning: a rapid, high resolution approach for spatial mechanical property mapping. Nanotechnology 22, 175707 (2011).
- J.L. Hutter and J. Bechhoefer, Calibration of atomic force microscope tips. Review of Scientific Instruments. 64, 1868–73 (1993).

This information has been sourced, reviewed and adapted from materials provided by Asylum Research - An Oxford Instruments Company.
For more information on this source, please visit Asylum Research - An Oxford Instruments Company.