Today, Carl Zeiss launches ATLAS™, a powerful hard- and software package, which, in combination with any scanning electron microscope from Carl Zeiss, enables quick and efficient imaging of large-area specimens with nanometer resolution.
ATLAS is initially being utilized both in the area of neurological research, e.g. the young field of brain mapping, and for traditional routine tasks in histology and pathology. Here, there is an increasing demand for efficient, cost-effective methods of examining a steadily rising number of specimens with constantly increasing sizes using resolutions in the nanometer range. There as well as in numerous future applications, ATLAS will offer users a new degree of productivity.
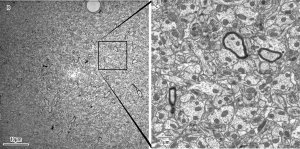 A 49 µm x 49 µm STEM-in-SEM image of rat hippocampus acquired on a ZEISS Supra 40 equipped with ATLAS™ (left) at 2 nm per pixel. The inset square covers a region of approximately 9 µm x 9 µm (right).
Courtesy of: John Mendenhall, Center for Learning and Memory, University of Texas at Austin
A 49 µm x 49 µm STEM-in-SEM image of rat hippocampus acquired on a ZEISS Supra 40 equipped with ATLAS™ (left) at 2 nm per pixel. The inset square covers a region of approximately 9 µm x 9 µm (right).
Courtesy of: John Mendenhall, Center for Learning and Memory, University of Texas at Austin
With suitable specimens, unattended operation can acquire multi-image montages that span extremely large fields of view, permitting capture of regions on the millimeter scale with resolution on the nanometer scale in a handful of hours. The in-built viewer software with integrated zoom function facilitates continous enlargement of the final image from rough overview until nanometer resolution.
The heart of the ATLAS system is an adaptive 16-bit scan generator and dual supersampling signal acquisition system, tightly integrated into the SmartSEM software for microscope control. ATLAS enables acquisition of individual images up to one gigapixel in size (32k x 32k), at up to sixteen bit pixel depth, and calls upon the rich suite of SmartSEM microscope automation features to allow automated acquisition of one or more image montages that may exceed one terapixel in size