Measuring small, poorly scattering molecules in solution has always been a challenge for dynamic light scattering systems. Getting the ultimate sensitivity in the past has required a specialized optical set-up dedicated to this type of measurement, or a powerful laser source which is unpopular because of its complexity and cost. This application note shows how the Malvern Panalytical HPPS can reach these sensitivities and maintain a wide range of applications.
Lysozyme
Lysozyme is an enzyme found in human tears and saliva, and is responsible for killing bacterial cells. It was discovered by Alexander Fleming in 1922 and became known as the first antibiotic.
It was the first protein to have its structure determined, as was found to be compact and nearly spherical. This fact along with its small size and easy availability makes lysozyme an excellent choice for investigating the sensitivity of a dynamic light scattering system.
Equipment Used
Malvern Panalytical HPPS.
Lysozyme (MERCK), 0.1 molar Sodium-Acetate buffer (pH 4.25), disposable plastic syringe, 0.2 μm syringe filter, square glass cuvette.
Sample Preparation and Measurement
Although only 15 μg of sample is required for a measurement, to enable an accurate concentration to be prepared, 100 mg Lysozyme was dissolved in 1000 ml Sodium-Acetate buffer. 0.2 ml was filtered via a 0.2 μm syringe filter into a clean, dust free glass cuvette. This constitutes a Lysozyme monomer concentration of 0.1mg/ml.
The glass cuvette was inserted into the unit and the temperature allowed to equilibrate for 3 minutes before a 300s measurement was performed.
Results
Size
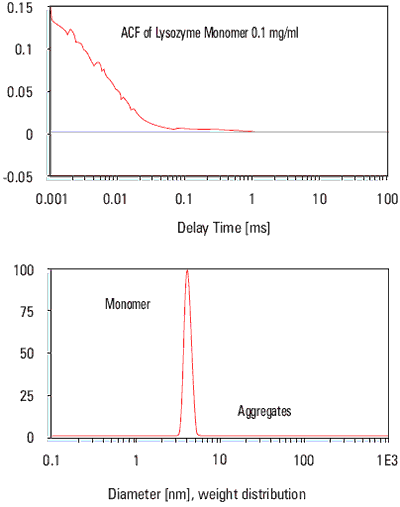
The correlation function in the diagram below indicates that the data is good enough to calculate the mean size and polydispersity of the Lysozyme, and an indication of the size distribution is possible.
|
|
|
Figure 1. Particle size distribution of the lysozyme sample.
|
Despite the low concentration, the monomer diameter of 4 nm was clearly measured with excellent precision. In addition, the aggregates, which are extremely difficult to avoid with such small and dilute samples, were measurable as well, the mass contribution of these aggregates was measured to be less then 0.3%.
This indicates that it is feasible to measure the size of 15 μg of Lysozyme.
Confirmation Of The Result Using The Scattering Intensity
The sample concentration of 0.1 mg/ml can be confirmed, and by inference the result validated by comparing this concentration with the value calculated from the scattering intensity of the sample. The detected photon count rate of the solution was measured as 29 kcps (Thousands of counts per second). The scattering from the dispersant was measured as 20.1 kcps, which gives 8.9 kcps as the scattering due to the Lysozyme monomer. We can calibrate the scattering intensity with a standard of known Rayleigh Ratio, in this case toluene, which gave a count rate of 220 kcps. Taking the dn/dc for Lysozyme as 0.2 at a wavelength of 632.8 nm, and a monomer Mw of 14,400 Dalton, the concentration was calculated to be 0.102 mg/ml, which was well within the weighing accuracy of 5% in this procedure.
Conclusion
The Malvern Panalytical HPPS is perfectly suited for DLS measurements on small particles even at very low concentration, as demonstrated here. Despite the low count rate, the full particle size range from the nm to the μm regime is accessible in a single measurement, which ensures that the instrument can be used to measure the monomer size and screen for aggregates.
.png)
This information has been sourced, reviewed and adapted from materials provided by Malvern Panalytical.
For more information please visit Malvern Panalytical.