DOI :
10.2240/azojono0118
Nov 8 2007
Table of Contents
Abstract
Background
Material and Methods
Results
Electronic Coupling
Discussion
Conclusion
Acknowledgement
References
Contact Details
Abstract
In the present manuscript, current-voltage (I-V) measurements of double stranded guanine rich sequences of λ-DNA have been reported. These sequences show length dependent conductivity. Conductivity (σ0) DNA of length, L = 0.6494 x 10-4 (1910 bp), 1 x 10-4 (2947 bp) and 1.3498 x 10-4 cm (3970 bp) was found to be 2.4 x 105, 7.7 x 102 Ω-1cm-1 and insulator behavior, respectively. I-V characterization of immobilized DNA was done on gold micro-electrodes fabricated by laser ablation using Optical Tweezer of power 0.66 mW. Charge transfer distance evaluation of intrinsic guanine rich sequences show increasing frequency of intervening bases between conducting units with increase in length. Outcomes of the present study can be useful for ascertaining the behavior of nanowires with varying charge transfer distance
Background
Biological and physical studies on DNA structure have revealed considerable interest into the electronic properties of DNA (1). Formation of DNA lesions as a result of radiation effect and ionization potential studies of nitrogenous bases of DNA has tracked researchers towards DNA electronics (2). Besides possessing requisite π-electron rich bases for conductivity, DNA also possesses nano-scale dimensions for nano-electronics (3, 4). These properties make DNA a promising material for molecular electronics. Electrical properties of DNA are being studied with the aim of producing nanoscale devices such as molecular wire (5, 6).
Earlier I-V characterization was implicated by photoinduced charge transfer studies (7), whereas recent studies have centered on direct electrical measurements. The electrical conductivity measurements have yielded ambiguous both experimental and theoretical results, pronouncing DNA possesses a wide range of behaviors. Researchers studied I-V characteristics of complete genome of λ-bacteriophage and chemically synthesized oligos up to 30 bp (8, 9). I-V characterization of natural DNA brings together low and high guanine rich regions separated by intervening sequences. Sequences with rich guanine content has high potential for future nanowire, as guanine is having lowest oxidation potential among four bases constituting DNA sequence (10). No more literature has been reported for I-V characteristics of guanine rich intrinsic sequences of λ-bacteriophage.
This study report on direct I-V measurements of double stranded intrinsic guanine rich λ-DNA sequences. Three guanine rich regions were selected for this study so that electrical behavior of DNA nanowire should not be affected as a result of low guanine content. These intrinsic sequences of different size were synthesized by Polymerase Chain Reaction (PCR) using specific thiolated primers.
Material and Methods
Specific Thiolated (5’ ends) primers Pr1 (1F-ATGCTTGAACCCGCCTATGC, 1R-TCACTTCATGCTTCGGCTTGAC), Pr2 (2F-TGGGATATTACGTCAGCGAGGAC, 2R-CACTTCATGCTTCGGCTTGAC) and Pr3 (3F-TGACTGCTGCTGCATTGACG, 3R-GCCATGATTACGCCAGTTGTAC) were decided on Gene runner 3.05 programme by plotting GC% vs. λ-DNA sequence (48502 bp). Primers were synthesized and procured from Bio Basics Inc., Canada. Standardization of amplification conditions for the specific primers (Pr1, Pr2 & Pr3) were done on MJ Research Gradient cycler (PTC-200). Different sets of conditions were used, so that maximum amplification can be obtained.
Optimum amplification conditions obtained were 560C, 560C and 550C and 1.5 mM Mg-+ ion concentrated. λ-DNA was used as template in order to amplify the fragment of 1910 bp (0.6494 x 10-4 cm), 2947 bp (1 x 10-4 cm) and 3970 bp (1.3498 x 10-4 cm). PCR synthesized guanine rich sequences (SQ1 = 1910 bp; SQ2 = 2947 bp; SQ3 = 3970 bp) were purified using Nucleotrap PCR Purification kit. A procedure to fabricate gold electrode with spacing 0.6, 1.0 and 1.3 μm had been described elsewhere (11). In brief, Microelectrodes were fabricated on gold coated (30 nm) glass wafer using Optical Tweezer Cum Microlaser Dissection Combi System. Gold was ablated by applying UV-laser (λ-337 nm) of 4 ns pulse duration with energy of 20 µJ and average power of 0.66 mW. Electrodes were cleaned with piranha solution [H2SO4:H2O2 : 3:1 (v/v)] as mentioned earlier (12).
A drop (0.2 μl) of prepared DNA samples (SQ1, SQ2 & SQ3) was pipetted out over physically separated electrodes for their immobilization so as to establish inter-element wiring. It was incubated for a period of 16 hours and then washed thoroughly with deionized distilled water. Eventually, it was nitrogen dried and I-V characterized on desktop probe station by signatone attached with the Hewlett-Packard HP4155A, Semiconductor Parameter Analyzer having an internal resistance of ≥1013 W and current resolution of 10 fA. λ-DNA was immobilized as mentioned earlier (12). All chemicals and enzymes of molecular biology grade were procured from Q.BIO gene; BIO BASIC INC., Canada; USBIOLOGICAL, USA; Sigma Aldrich, USA and PIERCE, USA. All the solutions were prepared in deionized (18 MΩ) distilled ultra pure water (ELGA Purelab ultra system). Samples were prepared in deionized water to exclude the role of counter ion effect in DNA conductance. Magnesium and other biological inorganic ions were not added as they unstuck bases of DNA and RNA (13).
Results
I-V Measurements: PCR synthesized guanine-rich (SQ1, SQ2 and SQ3) and λ-DNA sequences were immobilized between micro electrodes. The amount of DNA between electrodes was estimated to be ~ 1-4 x 10-1 ng for SQ1, SQ2 and SQ3 and ~ 4.0 x 10-2 ng for λ-DNA. Isotropic electrical characteristic was observed in each case. Hence, the structure of DNA is thought to be in amorphous state i.e. randomly distributed (14). To ensure that the conductivity observed was due to DNA and not because of any contamination, controlled experiments were performed. Electrodes with immobilized DNA were incubated for 30 minutes in a solution containing DNase-I.
This enzyme specifically cuts double stranded DNA. I-V characterization of DNase treated electrode shows no current. This ensures presence of DNA between electrodes. In another control experiment, electrodes were given microlaser treatment instead of DNase, again the result was same. The kind of characteristics obtained rule out the role of counter ions effects. It is not expected to result in sharp fall in conduction if there is a counter ions effect (Fig. 1D). The most accepted factor may contribute to conductivity along the DNA double helix is due to mobile counter ion of water thin film. While mobile counter ions may contribute to the conductivity at room temperature, the nitrogen drying of samples before carrying out conductivity measurement and the sharp fall down in conductivity with increasing length rules out for its significant role.
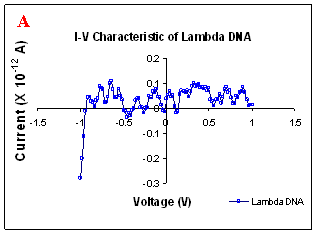
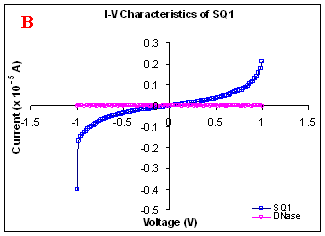
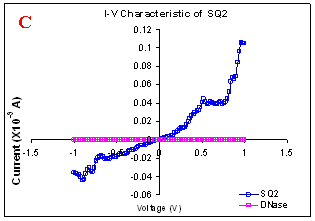
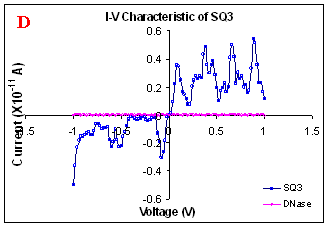 Figure 1. I-V characteristics of intrinsic sequences of λ-DNA. A- λ-DNA, B-SQ1, C-SQ2 and D-SQ3.
Figure 1. I-V characteristics of intrinsic sequences of λ-DNA. A- λ-DNA, B-SQ1, C-SQ2 and D-SQ3.
Figure 1 compares I-V characteristics of λ-DNA, SQ1, SQ2 and SQ3 at -1 to +1V. Current was measured at normal and reverse polarity (N/R), isotropic characteristics were observed. Voltage sweeps were performed both in negative to positive direction and the fine structure as well as the overall shape of the data is mirrored around zero bias for up compared to down sweeps. Average of three measurements for each case is shown in Figure 1 and the evaluation has been reported in Table 1 for SQ1, SQ2 and SQ3.
Table 1. I-V measurement conditions and outcomes for SQ1, SQ2 and SQ3.
|
|
|
SQ1
(1910 bp)
|
-1 to +1
|
0.6 μm
|
N/R
|
~ 0.16 eV
|
6.20 x 10-5 A
|
1.61 x 104 Ω
|
|
SQ2
(2947 bp)
|
-1 to +1
|
1.0 μm
|
N/R
|
~ 0.22 eV
|
3.23 x 10-8 A
|
3.09x 107 Ω
|
|
SQ3
(3970 bp)
|
-1 to +1
|
1.3 μm
|
N/R
|
~ 0.02 eV
|
1.37 x 10-10 A
|
7.29 x 109 Ω
|
Very low current in the range of pA was observed for λ-DNA (Fig. 1A). SQ1 and SQ2 are pronouncedly nonlinear, with a band gap up to ~0.16 and ~0.22 eV, beyond which sizable current flow occurs (Fig 1B & C). Current range of 10 μA and 10 nA was observed for SQ1 and SQ2, respectively. SQ3 shows almost similar behavior as in case of λ-DNA with a band gap of 0.02 eV and current in 10 pA range (Fig. 1D).
Electronic Coupling
Electronic coupling energy is an important ingredient for all the models describing DNA conductivity. Presently, it was calculated using single point calculations of the neutral G:C(A:T)nG:C on the B3LYP/6-31G(d, p) geometry using semi empirical intermediate neglect of differential overlap (INDO) Hamiltonian. The distance (r) between the base pairs i.e. distance between double membered aromatic moieties in third dimension was kept constant as in case of B-DNA is 3.38 Å. Eventually, electronic coupling energies for hole transfer were obtained from the energies of HOMO, and HOMO-1 of the stack of base pairs, obtained with the INDO Hamiltonian at the DFT/B3LYP optimized geometry (15, 16). Geometries of bases and base pairs in B-DNA were created using the templates for nucleic acids from the AMBER force field as implemented in HYPERCHEM. The sugar-phosphate backbone was removed and hydrogen was added at standard bond lengths. Base pair distance and the angle between the planes of two base pairs was kept 3.38 Å and 360, respectively.
Discussion
Very low current in pA range was observed for λ-DNA. This can be attributed to low threshold voltage, as the significant value of current was incurred for SQ1 and SQ2 at -1 to +1 V. Current in the range of 10 μA and 10 nA was shown by SQ1 and SQ2, respectively. At very first instant, this observation may be due to guanine richness of the sequences as the sequences selected were of guanine rich regions. On the other hand SQ3 has shown very dissimilar range of current i.e.10 pA. This current range is very close to current range observed for λ-DNA (Fig.1D). Even though, band gap was less for SQ3 as compared to SQ1 and SQ2 (Table1), sizable current was not observed. Percentage guanine in three intrinsic sequences of λ-DNA is ~ 58% and in λ-DNA is ~ 49%. GC percentage was calculated with respect to perspective length of DNA sequences used. Earlier, it was suggested (17) that insertion of a GC step into the otherwise A:T bridge actually decreases the efficiency of charge transport, which provides clear evidence that strict guanine hopping cannot describe long-range DNA-mediated charge transport.
Thus, it is the sequence length and intervening bases between conducting units ascertaining conductivity rather than guanine. However, the imperative role of guanine bases cannot be omitted. It is interesting to note that current range has diminished by a factor of 103 with consecutively increase in sequence length. Conductivity (σ0) evaluated for SQ1 and SQ2 with band gap of Δ=0.16 and Δ= 0.22 eV was found to be 2.4x105 and 7.7x102 Ω-1cm-1, respectively. To ascertain the comparative conductivity at the same band gap between SQ1 and SQ2, conductivity was calculated at Δ=0.16 eV for SQ2. It was calculated to be σ0 = 2.3x102 Ω-1cm-1. It does not show any significant difference from conductivity calculated at band gap of Δ= 0.22 eV. The maximum impedance [max (Imp)] offered by DNA segments SQ1, SQ2 and SQ3 were a factor of x104, x107 and x109, respectively. Conductivity was not evaluated for SQ3 as it has shown insulating behavior and high resistance of 109 Ω.
In order to ascertain the frequency of charge transfer distance in SQ1, SQ2 and SQ3 sequences, their analysis was done. It was observed that conducting units (GC: CG) are stepped in by (A:T or T:A)n bases [ G:C(A:T)nG:C]. Moreover, it was also found that ‘n’ varies from 1 to 10 with varying frequency in SQ1, SQ2 and SQ3. Frequency of n < 6 was found to be more as compared to frequency of n > 6. Stepping in tendency of ‘A’ or ‘T’ between conducting units ‘G’ was increased consecutively with increase in length.
To ascertain the role of intervening bases in DNA conductivity, electronic coupling energies were calculated for different frequency of intervening bases between conducting units (Table 3). It shows that electronic coupling energy is increasing with increase in number of A:T pairs between conducting units. Electronic coupling decreases sharply upto n = 3 and further increase in ‘n’ sharp fall is not observed. This results in weak coupling for two adjacent conducting units causing decrease in conductivity. The study and discussion about the charge carriers and the effect of nucleobase electronic coupling may be insufficient to draw firm conclusions about the charge transfer distance but it serves the purpose that intervening bases between conducting units causes significant change in electronic coupling energy which is a necessary ingredient for charge transfer.
Table 2. Coupling energies calculated for conducting units (G) with intervening bases (A).
|
|
|
GAG
|
-8.539
|
-8.488
|
0.051
|
|
G(A)2G
|
-8.564
|
-8.465
|
0.099
|
|
G(A)3G
|
-8.572
|
-8.455
|
0.117
|
|
G(A)4G
|
-8.575
|
-8.450
|
0.125
|
|
G(A)5G
|
-8.576
|
-8.448
|
0.128
|
|
G(A)6G
|
-8.576
|
-8.447
|
0.128
|
|
G(A)7G
|
-8.576
|
-8.444
|
0.132
|
|
G(A)8G
|
-8.576
|
-8.443
|
0.133
|
|
G(A)9G
|
-8.576
|
-8.442
|
0.134
|
|
G(A)10G
|
-4.938
|
-4.804
|
0.134
|
It is also reported that increasing number of (A:T)n (n>4) does not block the charge to move, rather A:T acts as a charge carrier (18). Table 3 also shows with increase in ‘n’ from 1-3 electronic coupling energy increases sharply and after n = 4 such trend diminishes. This result can be further corroborated by the studies of Saito et al. (1998) who calculated ionization energy for G:C is 7.34 eV, for TAGAT is 6.73 eV, for TTGTT is 6.96 eV (19). This clearly indicates that adjacent bases promote stabilization hence decrease in Ip for TAGAT and TTGTT. This accounts averaging of intervening sequences necessary for long distance conductivity. However, it is not necessary that every possible sequence differences will result in same pattern of conductivity.
Conclusion
In the present case, conductivity of intrinsic guanine rich sequences of λ-DNA was found to be length dependent. It is inferred from the sequences evaluation and electronic coupling energies calculation between two conducting units that conductivity of studied sequences was modified by frequency of intervening bases. Number of intervening bases between two conducting unit is not constant or fixed. Variability of intervening bases was found to be increasing with increase in DNA sequence length. DNA conductivity is not completely governed by guanine bases but also complemented by ‘AT’ bases. Averaging of intervening sequences is necessary for long distance charge transfer. These results may provide insights into the electrical behavior of guanine rich sequences with varying intervening bases. It may also be helpful in modifying the conductivity of DNA nanowire.
Acknowledgement
This work was supported by Department of Biotechnology (DBT) and Department of Science and Technology (DST). Authors are thankful to Dr. Prakash from GETECH Hyderabad, India for microelectrode array fabrication. We are also thankful to Mr. A.K. Shukla and Dr. Amit Sharma for their valuable guidance and suggestions. One of us authors (Ram Ajore) thanks Council of Scientific and Industrial Research (CSIR), Delhi for providing fellowship
References
- Bhalla V., Bajpai R.P. and Bharadwaj L.M., “DNA electronics”, EMBO reports, 4, 1-4, 2003.
- O'Neill P. and Fielden E.M., “Primary free radical processes in DNA”, Adv Radiat Biol, 17, 53, 1993.
- Warman J.M., de Hass M. P. and Rupprecht A., “DNA: A molecular wire”, Chemical Physical Letters, 294, 319-322, 1996.
- Joachim C., Gimzewski J.K. and Aviram A., “Electronics using hybrid- molecular and mono-molecular devices” Nature, 408, 541-548, 2000.
- Kasumov A.Y., Kodak M., Gueron S., Reulet B., Volkov V.T., Klinov D.V. and Bouchia H., “Proximity-induced superconductivity in DNA”, Science, 291, 280, 2001.
- Storm A.J., Noort J.V., Vries S.D. and Dekker C., “Insulating behavior for DNA molecules between nanoelectrodes at the 100 nm length scale”, Applied Physics Letters, 79, 3881-3883, 2001.
- Kelley S.O. and Barton J.K., “Electron transfer between bases in double helical DNA”, Science, 283, 375, 2002.
- Yoo K. -H., Ha D. H., Lee J. -O., Park J. W., Kim J., Kim J. J., Lee H. -Y., Kawai T. and Choi H. -Y., “Electrical conduction through poly(dA)-poly(dT) and poly(dG)-(dC) DNA molecules”, Phys. Rev. Lett, 87, 198102-05, 2001.
- Porath D., Bezryadin A., De Vries S. and Dekker C., “Direct measurement of electrical transport through DNA molecules”, Nature, 403, 635-638, 2000.
- Bharadwaj L.M., Kaur I., Kumar R. and Bajpai R.P., “Design simulation of DNA-based electronic components”, Proc. of SPIE, 4937, 226-230, 2002.
- Kumar S., Kumar R., Shukla A.K. and Bharadwaj L. M., “Microelectrodes fabrication using laser scissor”, Materials Letters, 61, 3829-3832, 2007.
- Ajore R., Kumar R., Kaur I., Sobti R.C. and Bharadwaj L.M., “DNA immobilization chemical interference due to aggregates study by dip and drop approach”, Journal of biochemical and biophysical methods, 70, 779-785, 2007.
- McFail-Isom L., Shui L. and Williams L D., “Divalent cations stabilize unstuck conformation of DNA AND RNA by interacting with base π system”, Biochemistry, 37, 17105-17111, 1998.
- Otsuka Y., Lee H.-Y., Gu J.-H., Lee J.-O., Yoo K.-H., Tanaka H., Tabata H. and Kawai T., “Influence of humidity on the electrical conductivity of synthesized DNA film on nanogap electrode”, Jpn. J. Appl. Phys, 41, 891-894, 2002.
- Lu S.-Z., Li X.-Y. and Liu J.-F., “Molecular Orbital Analysis in Evaluation of Electron-Transfer Matrix Element by Koopmans' Theory”, J. Phys. Chem A, 108, 4125, 2004.
- Li X.-Y., Tang X.-S. and He F.-C., “Electron transfer in poly(p-phenylene) oligomers: effect of external electric field and application of Koopmans theorem”, Chem. Phys, 248, 137, 1999.
- Williams T.T., Odom D.T. and Barton J.K., “Variation in DNA charge transport with nucleotide composition and sequence”, JACS, 122, 9048, 2000.
- Giese B. and Spitchy M., “Long distance charge transport through DNA: quantification and extension of the hopping model”.,Chem. Phys. Chem, 1,195, 2000.
- Saito I., Nakamura T., Nakatani K., Yoshioka Y., Yamaguchi K. and Sugiyama H., “Mapping of the Hot Spots for DNA damage by one-electron oxidation: Efficacy of GG doublets and GGG triplets as a Trap in long-range hole migration”, JACS, 120, 12686-12687, 1998.
Contact Details
Ram Ajore, Inderpreet Kaur, Lalit M. Bharadwaj
Biomolecular Electronics and Nanotechnology Division (BEND)
Central Scientific Instruments Organization (CSIO)
Sector-30C, Chandigarh
India
Phone: +91-172-2657811 Ext. 482, 452
+91-172-2656285
Fax: +91-172-2657267
E-mail: [email protected], [email protected]
R.C.Sobti
Department of Biotechnology, Panjab University
Sector-14, Chandigarh
India