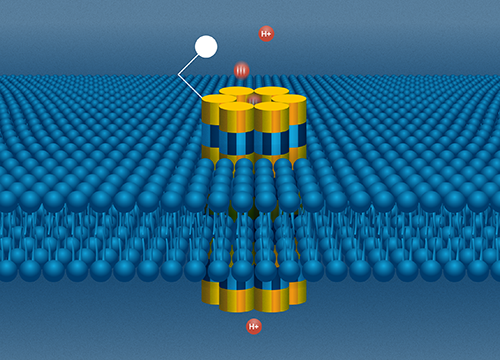
The bioprotonic system. A DNA nanopore sits in a lipid bilayer as an electrode sends a stream of protons through the channel. Image Credit: Molly Fine
The researchers can identify biomolecules that signal the presence of human disease using their bioprotonic system, a device that merges electronic components with biological components and employs electrical currents of protons, among other applications. The device’s specifications were just published in the journal Nature Communications.
Cells tend to be interconnected—they talk to each other, or they talk to the external environment, using these intermembrane channels. What we set out to do with our collaborators at MIT was create an artificial ion channel in a way that we could tune the properties of the ion channel and its functionality as we desire.
Marco Rolandi, Associate Professor, Electrical Engineering, University of California, Santa Cruz
The researchers at MIT can bioengineer a strand of DNA, which naturally takes the shape of a double helix, into whatever shape they choose using a process known as DNA origami. For this research, they developed a miniature tunnel precisely designed for a stream of protons (H-plus) to go through best. This small channel is known as a nanopore, and it was first proposed at UCSC.
Rolandi’s bioprotonic system, which is meant to simulate the watery, ion-conducting world of the cellular environment, houses the DNA nanopore. A double layer of lipids, analogous to a cell membrane, divides water, which represents the environment outside of the cell, from an electrode, which represents the inside of the cell, and the implanted nanopore works as a conduit between the two sides.
The electrode creates a stream of protons that go down the nanopore channel to the opposite side of the nanopore, where a molecule binding site can be tailored to attract certain biomolecules of interest. If one of those molecules is in the water, it will cling to the nanopore’s one end and obstruct the flow of protons through the channel.
The technology converts the proton signal into an electrical signal that can be read by the researchers. The researchers know that a biomolecule is present when the equipment does not detect protons traveling through the channel.
The device also has two handles composed of cholesterol that are positioned across the lipid bilayer to promote proton conductivity across the nanopore channel.
Rolandi added, “The uniqueness of the approach is the combination of these proton-conducting devices with supportive lipid bilayers, and I believe we are the only groups that are working on those, with this dock design for the DNA nanopores. The novelty is both the integration of the device and the ability to sense using these DNA nanopores.”
The researchers demonstrate in their study that they can utilize the bioprotonic system to detect the biomolecule B-type natriuretic peptide, which is a sign of heart disease. This demonstrates the device’s potential for biomolecule identification in an in-vitro or clinical context.
The device could one day include many nanopores, each of which would be set up to detect a different type of biomolecule, according to the researchers’ predictions for the future.
Rolandi concluded, “It is definitely part of the attractiveness of the system—in the near future we could multiplex, so we could have a full suite of biosensors.”
Le (Dante) Luo, Yunjeong Park, and Jesse Sanchez, all from UCSC, worked on this study. Additionally involved in this effort were researchers from the University of Washington and the TOBB University of Economics and Technology in Ankara, Turkey. The National Science Foundation funded the research.
Journal Reference
Luo, L., et al. (2023) DNA nanopores as artificial membrane channels for bioprotonics. Nature Communications. doi:10.1038/s41467-023-40870-1