A standard method of making a virus involves mixing proteins with a nucleic acid such as RNA or DNA, and then shaking the mixture. This method can produce a virus in just a matter of seconds. Whilst this method may appear strange and unsettling it is an easy and simple method of virus production.
Influenza, which is the cause of flu, is a virus that spreads easily and is capable of quickly assembling in large amounts. The self-assembly of viruses such as influenza has continued to be a long-standing mystery, because this process not only occurs rapidly, but also takes place in exceptionally short timescales.
A better understanding of the virus self-assembly process could help in the development of new types of drugs, which could work by preventing the formation of viruses in the first place.
In order to determine the self-assembly mechanism Harvard researchers have developed a novel system that enables the rapid monitoring of individual genomic molecules and proteins as they combine together to form a virus. With the help of this system viruses of a nanometer size can possibly be tracked at sub-millisecond time scales.
Our goal is to understand how viruses manage to assemble spontaneously, so quickly and so robustly.
Yoav Lahini - Harvard
Greater understanding of this self-assembly process could help in the development of artificial nanomaterials that instantly combine together. Also, better insight about the intermediate stages of the assembly process could help scientists to explore, track and interfere with this unique process.
However, the size and speed of the virus assembly process has continued to impede its study. Though fluorescent microscopy is capable of detecting individual proteins, photons produced by the fluorescent chemical compound are too slow to capture the process as it occurs.
This can be compared to attempting to capture the movements of a hummingbird’s wing with a stop-motion camera. Whilst the camera is able to record this process important frames, which show exactly what is happen, will always be missing.
Harvard sees viruses in new light
Capsid proteins are nano-size particles that can be observed by the way they scatter light. The proteins scatter light elastically, producing a countless number of photons in a single instant that can be measured. Howver, the photons that are produced interact with the reflected light, dust particles and other imperfections present in the optical path. All of these factors impede the tracking of nano-sized particles.
In order to overcome this the researchers developed a new approach which uses the unique properties of optical fibers. The team developed a novel optical fiber, which features a nano-scale channel, relatively smaller than the wavelength of light, running through the core of the silica fiber.
The channel is filled with liquid-containing nanoparticles so that when light is directed through the core of the fiber, the nanoparticles are scattered within the channel and subsequently collected by a microscope placed over the fiber.
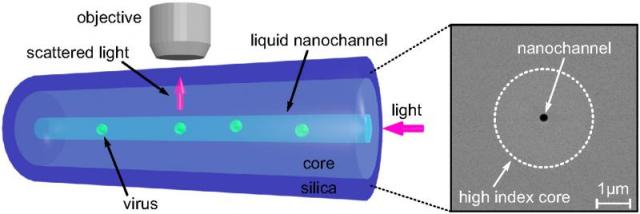
A schematic showing how the optical fiber works
The novel fiber-based method allowed the team to observe the movement of the viruses at a with thousands of measurements every second. The scientists are now planning to track individual viruses and viral proteins, which disperse significantly less light compared to a large virus sample.
These are the smallest viruses to be tracked on sub-millisecond time scales, which are comparable to the time scales for self-assembly.
Rees Garmann - Harvard
This research is a step forward in observing and measuring the self-assembly of viruses. Viral infection involves many complex molecular and cellular pathways, but self-assembly is a process that is found in many different viruses.
This simple technology, which is cheap, easy and scalable, could provide a new, cost effective way to study and diagnose viruses. From the point of view of fundamental physics, understanding the self-assembly of a naturally evolved system would be a major milestone in the study of complex systems.
Prof. Vinothan Manoharan - Harvard