Recent studies revealed that nanoparticles’ elasticity significantly influences nanoparticle physiological performance. For instance, it considerably affects nanoparticle biodistribution, systemic circulation lifetime, cellular uptake efficiency, and mode of cell internalization. Scientists stated that although it is clear that nanoparticle elasticity is an important physiological parameter that influences various biological processes, the underlying mechanism is not clear.
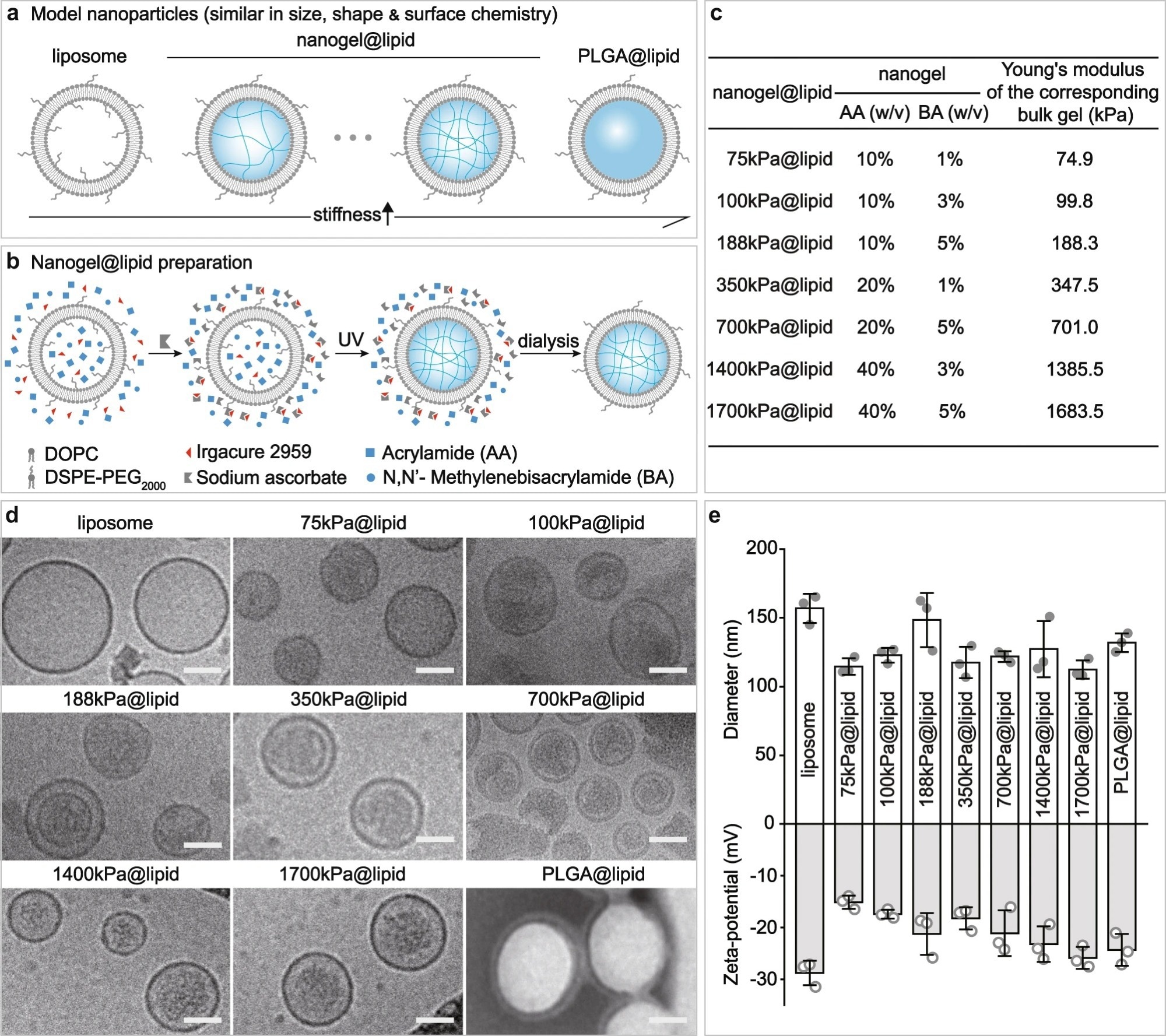
Figure 1. Our PEGylated model nanoparticles. a Schematic illustration on our model nanoparticles, which share a core-shell structure with a lipid bilayer shell of same composition while a core of tunable elasticity. b Schematic illustration on the preparation of a nanogel@lipid particle, whose elasticity can be tuned by adjusting its nanogel core’s crosslinking density. c Summary on the Young’s moduli of bulk gels which were prepared with corresponding hydrogel-precursor solutions used for preparing our nanogel@lipid particles. d Cryo-electron microscope (Cryo-EM) images of our liposome and nanogel@lipid nanoparticles, and transmission electron microscope (TEM) images of PLGA@lipid negatively stained with uranyl acetate. For each sample, microscopy images were taken in ≥5 different microscopy fields of view, and consistent results were observed. Scale bar = 50 nm. e Hydrodynamic diameters and surface zeta-potentials of our model nanoparticles. Bar heights are reported as average ± standard deviation (n = 3 independent experiments). © Li, M., Jin, X., Liu, T. et al. (2022)
When a nanoparticle is introduced into a living system, an immediate formation of protein corona occurs on the nanoparticle's surface by adsorbing proteins from surrounding biofluids. In this context, scientists have developed many strategies to prevent protein adsorption, including coating nanoparticles with polyethylene glycol (PEG). However, most strategies cannot fully eliminate the protein corona formation, but selectively restrict adherence of certain plasma proteins.
It is important to restrict the protein corona formation because it affects the functionalities of nanoparticles—for instance, the presence of protein corona on nanoparticles masks their targeting ability to specific ligands. Several studies have indicated that physiochemical properties of nanoparticles, such as their size, hydrophobicity, shape, surface change, etc., affect protein corona formation.
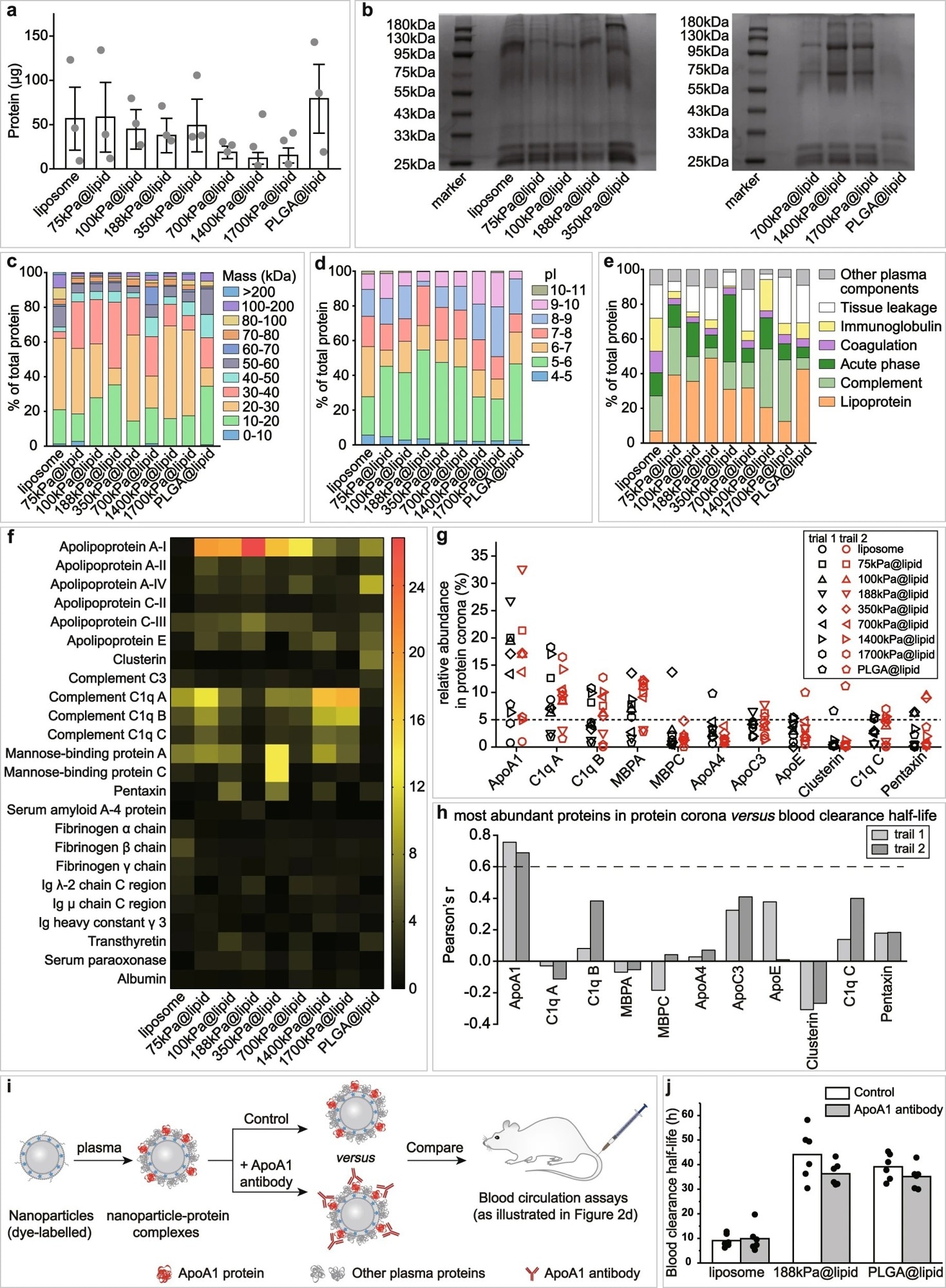
Figure 2. Effects of nanoparticle elasticity on protein corona. a Amounts of absorbed proteins on our nanoparticles (kept constant at 1 mg in lipid dose). Data points are reported as average ± standard deviation (n = 3 independent experiments). b Photographs of SDS-PAGE gels with absorbed proteins on our nanoparticles. The SDS-PAGE gel electrophoresis was only once. Classification of corona proteins according to c molecular weight, d calculated isoelectric point (pI), and e physiological functions. f Heat map of the most abundant proteins in protein coronas of our nanoparticles. g Distribution of the relative abundance in corona on different particles and h Pearson’s r between the relative abundance in corona and nanoparticle blood clearance half-life for proteins which exhibited a relative abundance of >5% on at least one nanoparticle. Pearson’s r of >0.6 and <−0.6 indicate strong positive and negative correlations, respectively. i Schematic illustration on blood circulation tests for examining the effects of pre-screening/pre-shielding the adsorbed ApoA1 in corona with ApoA1 antibody. j Blood clearance half-lives of 188 kPa@lipid with and without the adsorbed ApoA1 in corona pre-screened/pre-shielded by ApoA1 antibody, and those of liposome and PLGA@lipid in similar assays are included for reference. Bar heights are reported as average (n = 6 biologically independent mice). © Li, M., Jin, X., Liu, T. et al. (2022)
The Role of Nanoparticle Elasticity in Protein Corona Formation
Previous studies have reported that protein corona formation on nanoparticles of varying elasticity may lead to the production of different-sized nanoparticles. For instance, when the protein corona formation occurs on the surface of hard nanoparticles, an increase in the nanoparticle size is observed. However, liposomes are elastic and soft, resulting in differential surface coverage by specific proteins, which causes either an increase or decrease in the mean diameter of liposomes.
In the current study, scientists assessed if nanoparticle elasticity affects the formation of the protein corona. They further analyzed if certain plasma proteins affect nanoparticles’ elasticity-dependent fate. To study these aspects, model nanoparticles were used that helped isolate the effect of nanoparticle elasticity from other physiochemical parameters.
Researchers utilized core-shell nanoparticles as the model nanoparticle, similar to PEGylated lipid bilayer shell containing hydrogel nanoparticle cores of different crosslinking densities. In this model, the elasticity property of the hydrogel nanoparticle core was controlled by altering the crosslinking density, and the surface chemistry of the outer lipid bilayer shell was regulated by altering its composition.
Scientists studied the newly developed model nanoparticles using a transmission electron microscope (TEM) and cryo-electron microscopy (Cryo-EM). They reported that the model nanoparticles were spherical-shaped core-shelled structures.
In this study, the authors focus on the role of nanoparticle elasticity on cellular uptake efficiency and systemic circulation lifetime. In this context, they injected nanoparticles that were labeled with a membrane dye called DiD, into mice through the tail vein. They collected blood samples at particular time intervals and estimated nanoparticles’ blood clearance half-lives. The results contradict current understanding, as it revealed that softer nanoparticles do not always exhibit longer systemic circulation.
To understand how nanoparticle elasticity influences nanoparticle systematic circulation lifetime, scientists compiled a single plot that displayed the relationship between nanoparticle blood clearance half-life and nanoparticle elasticity. This plot showed a non-monotonic relationship that has been segmented into three distinct regions based on nanoparticle elasticity. Lower nanoparticle elasticity corresponds to longer blood clearance lifetimes. The plots developed to study the influence of nanoparticle elasticity on cellular uptake showed a lack of clear trends in this regard.
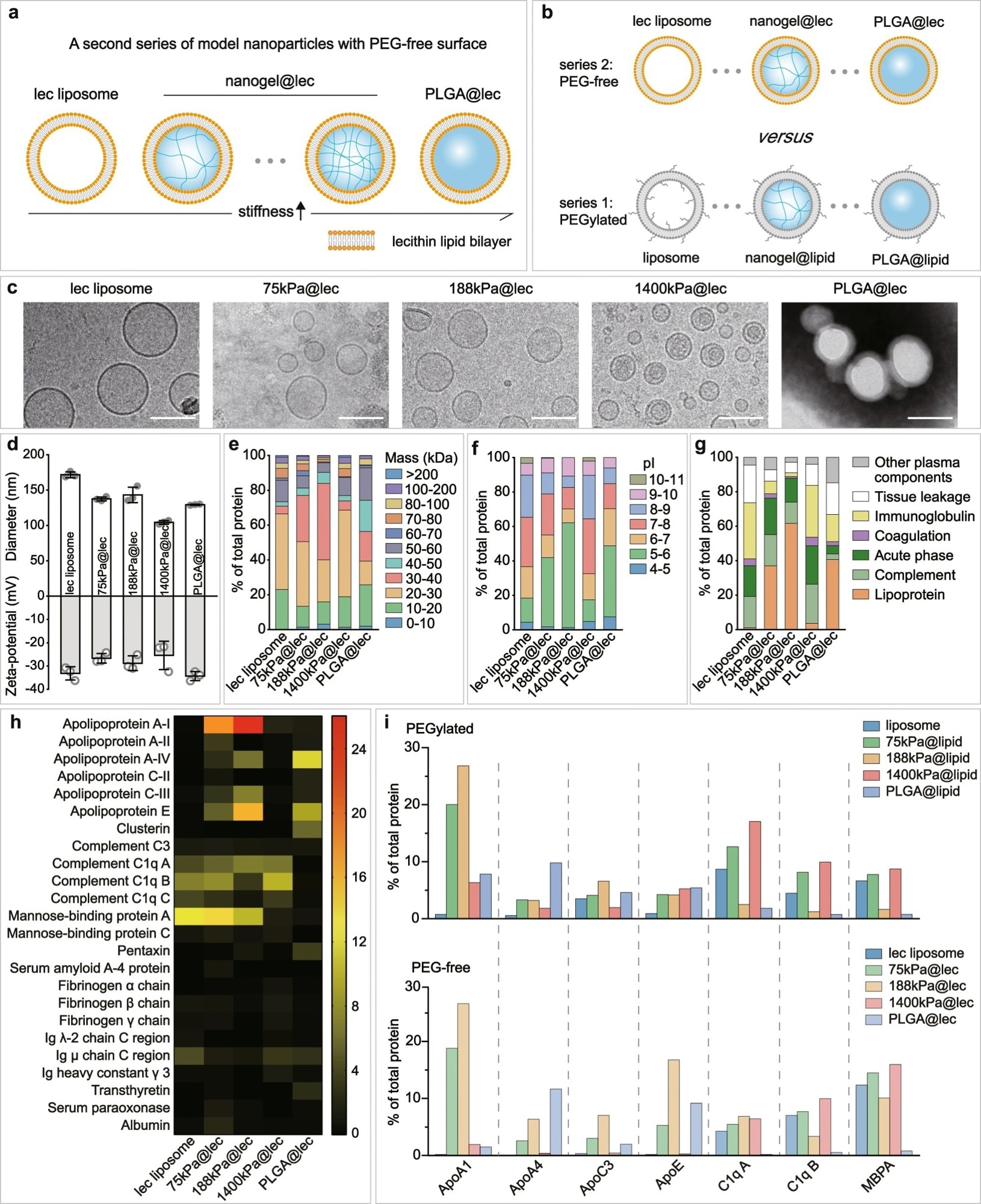
Figure 3. Nanoparticle elasticity overwhelms surface chemistry (PEG presence versus absence) in affecting protein corona. Schematic illustrations on a our PEG-free model nanoparticles, which have a core-shell structure with a lipid bilayer shell composed of lecithin (lec) and a core of tunable elasticity, and b a comparison with PEGylated nanoparticles. c Cryo-EM images of our lec liposome and nanogel@lec particles and TEM image of our PLGA@lec. For each sample, microscopy images were taken in ≥5 different microscopy fields of view, and consistent results were observed. Scale bar = 100 nm. d Hydrodynamic diameter and surface and zeta-potentials of our PEG-free nanoparticles. Bar heights are reported as average ± standard deviation (n = 3 independent experiments). Classifications of corona proteins according to e molecular weight, f calculated isoelectric point (pI), and g physiological functions for our PEG-free nanoparticles. h Heat map of the most abundant proteins in the coronas on our PEG-free nanoparticles. i Comparison on the distribution of the relative abundance in corona on our (top) PEGylated and (bottom) PEG-free nanoparticles for proteins which exhibited a relative abundance in corona of >5% on at least one nanoparticle. © Li, M., Jin, X., Liu, T. et al. (2022)
How Does Nanoparticle Elasticity Affect Protein Corona Formation?
Protein corona formed when model nanoparticles were incubated with mouse plasma. The composition of protein corona varied non-monotonically based on nanoparticle elasticity. Scientists reported apolipoprotein A-I (ApoA1) to be the corona protein that accumulated on the surface of nanoparticles, depending upon nanoparticle elasticity.
Compared to other plasma proteins, ApoA1 was found to be abundantly present, which positively correlated with nanoparticles’ systemic circulation lifetime in mice. Scientists revealed that ApoA1 could selectively bind to nanoparticles of intermediate elasticity (75–700 kPa) compared to softer or stiff nanoparticles.
To conclude, the authors demonstrated the mechanism by which nanoparticle elasticity affects nanoparticles’ systemic circulation lifetime. They reported that protein corona, i.e., ApoA1, is the key mechanism by which nanoparticle elasticity influences physiological properties. The current study indicates that nanoparticle elasticity is an easily tunable physiochemical parameter, which can be explored in the future to alter the formation of the protein corona.
Reference
Li, M., Jin, X., Liu, T. et al. (2022) Nanoparticle elasticity affects systemic circulation lifetime by modulating adsorption of apolipoprotein A-I in corona formation. Nature Communications, 13 (4137). https://doi.org/10.1038/s41467-022-31882-4
Disclaimer: The views expressed here are those of the author expressed in their private capacity and do not necessarily represent the views of AZoM.com Limited T/A AZoNetwork the owner and operator of this website. This disclaimer forms part of the Terms and conditions of use of this website.